I am interested in various extensions to the simple Bayesian model for accrual that Byron Gajewski and I derived and published in Statistics in Medicine. An important extension would be accrual in multi-center trials. A hierarchical model makes a lot of sense in this case, so I wanted to examine a simple hierarchical model that appears in the BUGS manual.
The particular example shows the number of failures for 10 mechanical pumps. A slight complication is that there was variation in the amount of time that each pump was monitored for failures. Here’s the raw data:
i x t
1 5 94.30
2 1 15.70
3 5 62.90
4 14 126.00
5 3 5.24
6 19 31.40
7 1 1.05
8 1 1.05
9 4 2.10
10 22 10.50
In this table, i represents the number of the pump, x represents the number of failures and t represents the monitoring time. Notice that there is substantial variation in the monitoring times.
You can get a crude failure rate by dividing x by t.
i x t r
1 5 94.30 0.053
2 1 15.70 0.064
3 5 62.90 0.079
4 14 126.00 0.111
5 3 5.24 0.573
6 19 31.40 0.605
7 1 1.05 0.952
8 1 1.05 0.952
9 4 2.10 1.905
10 22 10.50 2.095
The last column, r, represents the failure rate per unit time. Notice that there is a 40 fold difference between the lowest failure rate and the highest failure rate.
A hierarchical model would assume that each pump would have its own failure rate parameter, but that these parameters would be drawn from a common hyperdistribution. This is model is easy to fit in BUGS. Here’s the code for that model, taken directly from the BUGS manual.
model {
for (i in 1 : N) {
theta[i] ~ dgamma(alpha, beta)
lambda[i] <- theta[i] * t[i]
x[i] ~ dpois(lambda[i])
}
alpha ~ dexp(1)
beta ~ dgamma(0.1, 1.0)
}
Now I wanted to use R to fit the BUGS model, but as I noted in an earlier webpage, the BRugs libary does not work with recent versions of R. I switched to the rbugs library, and here is the code I used.
library("rbugs")
bugs.path <- "c:/Program Files (x86)/OpenBUGS/OpenBUGS321/OpenBUGS.exe"
bugs.wd <- "I:/tmp"
pumps.dat <- list(t = c(94.3, 15.7, 62.9, 126, 5.24, 31.4, 1.05, 1.05, 2.1, 10.5),
x = c( 5, 1, 5, 14, 3, 19, 1, 1, 4, 22), N = 10)
pumps.inits <- list(list(alpha = 1, beta = 1), list(alpha = 10, beta = 10))
pumps.model <- "PumpsModel.txt"
pumps.pars <- c("theta", "alpha", "beta")
pumps.sim <- rbugs(data = pumps.dat, inits = pumps.inits,
paramSet = pumps.pars, model = pumps.model,
n.chains = 2, n.iter = 10000, n.burnin = 1000, n.thin = 1,
bugs=bugs.path, workingDir = bugs.wd, bugsWorkingDir = bugs.wd,
dic = TRUE)
The rbugs library requires you to specify that full path for the BUGS program, and depending on your operating system, it might be different for your computer. The data for BUGS is usually stored as a list, and the sample size (N) is normally part of this list. You need to specify initial values for alpha and beta (thankfully not for the ten theta values) and since I wanted to look at two chains, I specified two lists within a bigger list.
The BUGS code (shown above) is not part of the R program in this example, but instead is a separate file. It’s a bit tricky to place the BUGS code directly in the R program.
When this program is completed, the BUGS output is stored in the pumps.sim object. The iterations in the Markov Chain Monte Carlo are stored as matrices with the names chain1 and chain2. Here’s the code I used to extract the relevant information and plot it.
pumps.comb <- rbind(pumps.sim$chain1, pumps.sim$chain2)
pumps.quant <- apply(pumps.comb,2,quantile,probs=c(0.025,0.5,0.975))
png(file="pumps%02d.png",width=400,height=300)
par(las=1,mar=c(5.1,5.1,0.1,0.1))
plot(c(0,4),c(1,10),type="n",xlab="theta",ylab=" ",axes=FALSE)
axis(side=1)
axis(side=2,at=1:10,labels=paste("Pump #",10:1,sep=""))
segments(pumps.quant["2.5%",4:13],10:1,pumps.quant["97.5%",4:13],10:1)
segments(pumps.quant["50%",4:13],(10:1)-0.25,pumps.quant["50%",4:13],(10:1)+0.25)
u <- seq(0.01,4,length=1000)
d <- dgamma(u,shape=pumps.quant["50%","alpha"],rate=pumps.quant["50%","beta"])
plot(u,d,type="l",axes=FALSE,xlab="theta",ylab="density")
axis(side=1)
axis(side=2)
The first graph produced by this code shows 95% credible intervals for the individual failure rates.
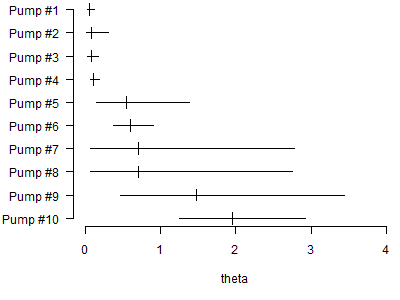
The second graph shows the hyper distribution for global failure rates.
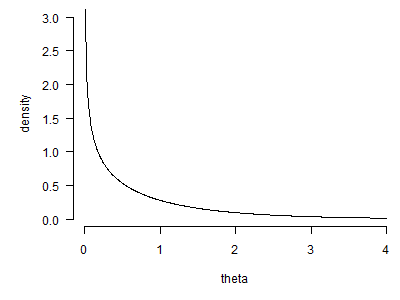
This looks like an exponential distribution, but actually it is even a bit more skewed than the exponential distribution. With one more line of code,
round(t(pumps.quant[,4:13]),2)
you can also produce a nice table of the credible intervals.
2.5% 50% 97.5%
theta[1] 0.02 0.06 0.12
theta[2] 0.01 0.08 0.30
theta[3] 0.03 0.08 0.18
theta[4] 0.06 0.11 0.18
theta[5] 0.15 0.54 1.38
theta[6] 0.37 0.60 0.91
theta[7] 0.07 0.71 2.77
theta[8] 0.07 0.70 2.76
theta[9] 0.47 1.47 3.45
theta[10] 1.25 1.96 2.92
It is trivial to run this example within BUGS, but running it through R allows you to create nice graphs and tables like these.
It is wothwhile to compare the crude rates to the rates estimated by BUGS. The rates estimated by BUGS show shrinkage towards the mean with greater shrinkage for those pumps with less data and for those pumps which have a rate further away from the overal rate.
You can find an earlier version of this page on my original website.